Telomerase reverse transcriptase promoter mutations in hepatocellular carcinogenesis
Abstract
Through several studies exploiting next-generation sequencing, we are obtaining a clearer picture of the complex genetic and molecular landscape of hepatocellular carcinoma (HCC). Consistent with the findings of other cancer types, telomerase reverse transcriptase (TERT) promoter mutations have been frequently reported in HCC. C228T and C250T are two major types of hot spot mutations in the TERT promoter region. Besides, in hepatitis B virus (HBV)-related HCC cases, the TERT promoter is recurrently interrupted by integration of HBV DNA. TERT promoter mutations are thought to be an early event in HCC carcinogenesis, and they are significantly associated with disease progression. In this review, we provide an updated overview of the somatic mutations in the TERT promoter region and discuss their possible roles in the development of HCC.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is one of the most common and deadliest cancers worldwide, ranking fifth and ninth in incidence, and second and sixth in mortality for males and females, respectively[1,2]. So far, only three molecular targeted agents, including sorafenib, lenvatinib and regorafenib, have been approved by the Food and Drug Administration for the treatment of HCC[3,4], and they only extend median survival by a few weeks to months[5]. Therefore, more research is needed to fill the gaps in knowledge of the genetic and molecular landscape of HCC in order to develop target therapies. The genetic landscape of HCC is complicated and involves a number of pathways as well as a considerable amount of somatic mutations in a wide range of genes[6]. Among all these genetic alterations, telomerase reverse transcriptase (TERT) promoter mutations occur most frequently, affecting ~60% of all HCC patients[6-8]. In this mini-review, we mainly summarize the frequency, mechanisms and clinical prospect of TERT promoter mutations in HCC. To provide more background information, this review also briefly touches upon the TERT promoter mutations in various cancers, although HCC remains the main focus of our discussion throughout the whole paper.
The structure and function of TERT
Human telomerase is a ribonucleoprotein polymerase that reverses the continuous telomere shortening in cell division by adding 5’-TTAGGG-3’ repeats to the ends of chromosome[9]. It consists of two core subunits: the catalytic component TERT and the RNA component (TERC) that serves as a template for elongating telomeres[10,11].
The TERT component is encoded by the TERT gene, located on chromosome 5 in humans. It spans a length of about 40,000 base pairs (bp) with 16 exons[12]. Of note, the TERT gene is suppressed in most normal somatic cells (excluding germ cells and stem cells), ensuring that these cells only divide a finite number of times and do not surpass the Hayflick limit[13,14]. Normal somatic cells stop dividing when their telomeres become critically short, whereupon they enter a stage called senescence[15]. Cancer cells, however, overcome replicative senescence and achieve immortality by reactivating the TERT gene and upregulating TERT expression[14].
The regulation of TERT expression largely depends on the activity of the TERT promoter, especially the core functional fragment that consists of a 260 bp DNA sequence with several transcription factor binding sites, but distinctly lacking a TATA box or a similar sequence[16,17]. The binding motifs in the TERT promoter include two evolutionarily-conserved E-boxes (CACGTG), located at -242 bp and -34 bp to the translational start site, for c-Myc binding[18]. The binding of c-Myc to the E-box activates TERT transcription, suggesting a role of c-Myc in regulation of the expression of TERT[19,20]. GC-boxes (GGGCGG), the binding sites for zinc finger transcription factor Sp1, are the other characteristic sequences in the TERT promoter region[21]. There are at least five GC-boxes within the core promoter of TERT, and they function synergistically to maintain the promoter activity of TERT[22]. P53 has been shown to down-regulate TERT transcription in an SP1-dependent manner[23].
TERT promoter mutations in several cancers
TERT promoter mutations are the most frequent somatic mutations in a variety of cancers. It has been widely reported that the two most common types of recurrent TERT promoter mutations are C228T and C250T, located at positions 1,295,228 and 1,295,250 on chromosome 5, or -124 bp and -146 bp of the ATG translational start site of the TERT gene[24-27]. In a systematic analysis involving 1,581 cancer cases of different types, 27.0% were found to have TERT promoter mutations[25]. Killela et al.[28] examined 1,230 tumor specimens of 60 different types and identified 231 TERT promoter mutations (18.8% of the total), among which C228T and C250T mutations accounted for 98%. Similarly, in a study where 1,515 tumors of the central nervous system were tested, 327 (21.6%) had TERT promoter mutations, and all except two contained either C228T or C250T[29]. Another study examined 150 cell lines of several cancer types from the Cancer Cell Line Encyclopedia and noted that 24 cell lines (16%) harbored either C228T or C250T mutations[26]. Statistics show that C228T is somewhat more prevalent than the C250T mutation [Table 1] in a wide range of cancer types, including various subtypes of CNS cancers, urogenital cancers, melanoma and thyroid cancer[25,26,28-37].
Telomerase reverse transcriptase promoter mutations in multiple cancers
| Cancer type | Number of cancer cases | Number of TERT mutations* | Number of different types of TERT promoter mutations** | Methods | Ref. | ||
|---|---|---|---|---|---|---|---|
| C228T | C250T | C228T or C250T | |||||
| Cancer tissue | |||||||
| Glioma, medulloblastoma, hepatocellular carcinoma, etc. | 1230 | 231 (18.8) | 179 (77.5) | 48 (20.8) | 227 (98.3) | PCR/Sanger sequencing | [28] |
| Bladder cancer, liver cancer, glioma, etc. | 1581 | 426 (26.9) | / | / | / | Whole-genome/low-pass whole-genome sequencing | [25] |
| CNS cancers | 1515 | 327 (21.6) | 257 (78.6) | 68 (20.8) | 325 (99.4) | PCR/bidirectional sequencing | [29] |
| CNS, bladder, thyroid cancers, etc. | 741 | 142 (19.2) | 99 (69.6) | 43 (30.3) | 140 (98.6) | PCR/Sanger sequencing | [36] |
| Urogenital cancers | 302 | 130 (43.0) | 100 (76.9) | 24 (18.5) | 124 (96.4) | PCR/Sanger sequencing | [37] |
| Medulloblastoma | 466 | 98 (21.0) | / | / | / | PCR/Sanger sequencing | [35] |
| Melanoma | 287 | 109 (38.0) | 51 (46.8) | 40 (36.7) | 91 (83.5) | PCR/Sanger sequencing | [32] |
| Bladder cancer | 262 | 218 (83.2) | 165 (75.7) | 32 (14.7) | 197 (90.4) | SNaPshot assay and Sanger sequencing | [34] |
| Melanoma | 77 | 24 (31.2) | 7 (29.2) | 5 (20.8) | 12 (50.0) | High-throughput sequencing/Sanger sequencing | [33] |
| Cancer cell line | |||||||
| Melanoma | 168 | 125 (74.4) | 46 (36.8) | 64 (51.2) | 110(88) | High-throughput sequencing/Sanger sequencing | [33] |
| Melanoma, liver, bladder cancers, etc. | 150 | 24 (36.0) | / | / | 24(100) | Whole-genome sequencing, Sanger sequencing, | [26] |
| Urothelial bladder cancer | 23 | 20 (87.0) | 16 (80.0) | 2 (10.0) | 18 (90.0) | PCR/Sanger sequencing | [31] |
| Urothelial bladder cancer | 32 | 28 (87.5) | 25 (89.3) | 3 (10.7) | 28(100) | PCR/Sanger sequencing | [30] |
Overall, it is widely accepted that glioma, melanoma, bladder cancer and HCC are among those commonly-affected by TERT promoter mutations[25,28,38].
TERT promoter mutations in HCC
The genomic landscape of HCC involves a number of pathways as well as somatic mutations in a wide range of genes, including TP53, CTNNB1, AXIN1, CDKN2A, ARID2, ARID1A, TSC1/TSC2, RPS6KA3, KEAP1, MLL2, and several epigenetic modifications[6]. Despite the complexity of the genomic landscape of HCC, the single most significant factor is genomic changes on TERT promoter, which include point mutations, hepatitis B virus (HBV) DNA integrations, amplifications and epigenetic modifications. TERT promoter point mutations contribute more frequently (54%-60%) to the reactivation of telomerase in HCC than the exclusively-present HBV insertions in the TERT promoter (10%-15%) and TERT amplification (5%-6%)[6-8]. Therefore, we are going to thoroughly discuss TERT promoter mutations while briefly touching upon other genomic and epigenomic alterations on TERT promoter in HCC.
TERT promoter point mutations
A few prominent studies on HCC demonstrated that TERT promoter mutations were found in about 30%-60% of the total cases[8,39-49]. Consistent with the findings in other cancer types, the two most common mutations were C228T and C250T, and the former was more prevalent than the latter in HCC [Table 2][8,39-47]. As shown in Table 2, there are no cases with both C228T and C250T mutations, which implies that these two hot spot mutations are mutually exclusive. Furthermore, a comprehensive review evaluating the distribution of TERT promoter mutations in 1,939 primary HCC from four continents also showed that TERT promoter mutations had almost the same level of prevalence in all continents, with slightly higher mutation rates in Europe (56.6%) and Africa (53.3%) than in America (40%) and Asia (42.5%), and that C228T mutation was universally more frequent than C250T[1].
Telomerase reverse transcriptase promoter mutations in hepatocellular carcinoma
| Number of HCC cases | Number of TERT mutations (%) | Number of different types of TERT promoter mutations (%*) | Methods | Ref. | ||
|---|---|---|---|---|---|---|
| C228T | C250T | C250T or C228T | ||||
| 469 | 254 (54.2) | 236 (92.9) | 11 (4.3) | 247 (97.2) | PCR/bidirectional sequencing | [8] |
| 316 | 103 (32.6) | 96 (93.2) | 5 (4.9) | 101 (98.1) | PCR/Sanger sequencing | [43] |
| 305 | 179 (58.7) | 166 (92.7) | 11 (6.1) | 177 (98.9) | PCR/Sanger sequencing | [42] |
| 276 | 85 (30.8) | 84 (98.8) | 1 (1.2) | 85(100) | PCR/Sanger sequencing | [44] |
| 196 | 87 (44.4) | / | / | / | Whole-genome sequencing | [48] |
| 195 | 57 (29.5) | 54 (94.7) | 3 (5.3) | 57(100) | PCR/Sanger sequencing | [45] |
| 160 | 46 (28.8) | 32 (69.6) | 14 (30.4) | 46(100) | PCR/Sanger sequencing | [39] |
| 44 | 15 (34.1) | 10(66) | 5(34) | 15(100) | PCR/Sanger Sequencing | [40] |
| 190 | 57 (30.0) | 50 (87.7) | 7 (12.3) | 57(100) | PCR/bidirectional sequencing | [46] |
| 127 | 64 (50.4) | 62 (96.9) | 2 (3.2) | 64(100) | PCR/Sanger sequencing | [47] |
| 123 | 45 (36.6) | 43 (95.6) | 2 (4.4) | 45(100) | PCR/Sanger sequencing | [41] |
| 125 | 85 (68.0) | / | / | / | PCR/Sanger sequencing | [49] |
Apart from the high frequency of TERT promoter mutations in HCC, another piece of useful information indicated by several lines of evidence is that TERT promoter mutations are associated with a few factors, including virus status, gender, age and tumor size of the patients. TERT promoter mutations were more frequent in HCC patients infected with hepatitis C virus[7,8,39,41,42,47,48,50] than in those infected by HBV. One study suggested that this phenomenon could be explained by the high rate of HBV DNA insertions in the TERT promoter[42]. Furthermore, several studies reported higher TERT promoter mutations rate in men[7,39,42], in older patients[7,50], in patients with smoking[51], in patients with smaller tumors[42], in patients with low serum levels of alpha-fetoprotein[42], and in patients with CTNNB1 mutations[8,42,47], while other papers either disagreed with or did not find these associations.
Further, TERT promoter mutations are early somatic genetic alterations in hepatocarcinogenesis, playing important roles in malignant transformation of preneoplastic cirrhotic lesions[42,52]. Nault et al.[52] found that the frequency of TERT promoter mutations increased as premalignant lesions transformed into HCC, from 6% in low-grade dysplastic nodules and 19% in high-grade dysplastic nodules to 61% in early HCC and 42% in small and progressed HCC; mutations in 10 other recurrent genes only emerged in small and progressed HCC. Similarly, Huang et al.[43] demonstrated that the mutation rates also increased in a stepwise manner during advanced HCC progression and reached a maximum of 45% in patients with stage C. Calderaro et al.[53] found that there were 64.6% (208/322) cases with TERT promotor mutations; HCC phenotypes were tightly associated with gene mutations, including TERT promoter mutations, and transcriptomic classification.
As the proportion of nonalcoholic fatty liver disease (NAFLD)-related HCC patients is increasing due to increased prevalence of metabolic syndrome, especially in Western countries[54-56], there have been studies investigating TERT promoter mutations in NAFLD-related HCC. One research analyzed the genetic aberrations of 11 tumor samples from 10 NAFLD-HCC patients and found that TERT promoter mutation C228T occurred in 9/11 (82%) cases[56]. On the contrary, in another study, the prevalence of TERT promoter mutations C228T and C250T was very low (3.2%) in patients with NAFLD[57]. Obviously, the TERT promoter mutation state in NAFLD-related HCC is far from conclusive.
TERT promoter insertional mutations by HBV DNA integration
HBV infection has been shown to be a causative factor of HCC, especially in Asians where chronic hepatitis B infection is prevalent. Integration of HBV DNA into the human genome of HCC cells is evident in HBV-related HCC[8,40,48,58-64]. Several lines of evidence demonstrate that the integration sites of HBV are not random. Integration of certain genomic sites, including near or within the genes of TERT[8,48,59-65], MLL4[48,59,61-63,65] and CCNE1[48,61-63,65] are more frequently identified in HCC[48].
To date, 13 independent studies have identified a total of 262 integrations of HBV DNA in the TERT gene, meaning that in more than 20% HBV-related HCC cases, TERT gene is interrupted by HBV integration[7,58,65-75]. TERT is the most susceptible gene for HBV integration, followed by MLL4 (79 integrations), CCNE1 (22 integrations) and CCNA2 (19 integrations)[76]. According to our pool analysis of the results from these articles[7,58,65-75], among the 262 HBV integrations in TERT, 73.28% (192/262) occur in the TERT promoter region, including 26% in the core functional fragment (-223 bp to -14 bp from the ATG translational start site). As the regulation of TERT expression largely depends on the activity of the TERT promoter region, especially the core functional fragment, HBV integration in the TERT promoter may have an important functional role in HCC development.
A few studies suggested that HBV tended to integrate in common chromosomal fragile sites, where DNA replication was delayed and DNA sequences were more susceptible to breakage[63,64]. Nevertheless, the findings that TERT was a recurrent integration site but not a fragile site demand new explanation[64]. More recent studies have therefore presented new possibilities. One study proposed that HBV preferentially integrates into TERT gene because disruption at these loci lowers the threshold for malignant transformation and thus grants a selective advantage to carcinogenesis[59]. Another two studies, using a similar line of reasoning, suggested that the recurrence of HBV integrations into TERT promoter region in HCC could be due to the potential growth advantage that augmented TERT expression provides for the clonal expansion and carcinogenesis of hepatocytes[60,62]. In TCGA database, the HCC with HBV DNA insertion into the TERT promoter displays the highest level of TERT RNA expression among all HCCs, suggesting an HBV cis-activating event did exist[48].
HBV integrations promote the development of HCC by inducing global genomic instability, elevating expression of adjacent genes, viral-host fusion transcripts and secondary mutations of host or viral genes, as well as by DNA copy number variations and proteins with oncogenic activity (X and preS gene products)[58,61,64,65]. Recently, based on the discovery that both HBV integration and somatic mutations in the TERT promoter were more frequent in male patients with HCC, Li et al.[69] proposed a novel mechanism in which sex hormones, along with GABPA play a role in regulating TERT expression. They analyzed 101 HBV-related HCC cases using a capture-next-generation sequencing platform and concluded with convincing evidence that the integration of HBV DNA, whose sequence contains both androgen- and estrogen-responsive elements, into the TERT promoter permits the androgen-receptor to up-regulate and the estrogen-receptor to down-regulate TERT transcription in a HNF4α-dependent manner[62].
Other genomic and epigenomic alterations on TERT promoter in HCC
TERT amplification in HCC
Totoki et al.[8] showed that TERT focal amplification was detected in 6.7% of the total 608 cases. Schulze et al.[77] observed less than 5% of TERT focal amplification in the 243 liver tumors. However, while both studies described the occurrence of TERT focal amplification in HCC, none of them investigated its effect on TERT expression level. Thus, more research is needed to confirm the role of TERT amplification in liver carcinogenesis.
Epigenetic modification of TERT promoter in HCC
As for epigenetic regulation of TERT promoter in HCC, Iliopoulos et al.[78] observed a strong negative correlation between TERT promoter methylation and TERT expression in all liver tissues they studied, proposing for the first time that the hypermethylation of TERT promoter and the methylation of histone H3-K9 resulted in the inhibition of c-Myc binding in E-box 1, which in turn inactivated TERT expression. However, this result contrasts with previous studies, which showed that TERT promoter epigenetic modification had either a positive correlation or no correlation with TERT expression and telomerase activity in other cancer types[79-83]. A more recent study examining 125 HCC cases in the Han Chinese population found that the promoter of the TERT gene is significantly hypermethylated, and it further showed that the hypermethylation is associated with higher expression of TERT, suggesting that TERT promoter hypermethylation contributes to the progression of liver carcinogenesis via elevating TERT expression level[84]. Overall, there is no definite conclusion regarding whether hypermethylation of TERT promoter has a positive or negative correlation with TERT expression and telomerase activity.
Mechanisms of TERT promoter mutations contributing to the development of HCC and other cancers
Although TERT promoter mutations are strongly associated with several cancers, the mechanism by which TERT promoter mutations lead to cancer development is not fully understood. How TERT promoter mutations increase TERT expression and whether the up-regulation of TERT directly translates into active telomerase activity that eventually contributes to tumorigenesis are two important questions requiring answers.
Mechanisms of TERT promoter in other cancers
It is currently accepted that C228T and C250T, the two most common mutation types in TERT promoter region, both create an 11-bp binding motif (5’-CCCCTTCCGGG-3’) for E-twenty-six (ETS) transcription factors[26,85,86]. In glioblastoma, a total of five ETS transcription factors were found (ELF1, ETS1, ETV3, ETV4 and GABPA) that modulate TERT expression. GABPA complexes with GABPB to form a fully functional heterodimer GABP transactivator, it was the only factor that reproducibly regulated TERT expression in a mutation-specific manner[86]. Akincilar et al.[24], using cell lines from several cancer types, including melanoma, glioblastoma, colon, and prostate cancers, etc., reported that TERT promoter mutations enhanced the binding of GABPA, mediating long-range chromatin interaction (at chr5: 1,556,087-1,558,758, a region 300 kb upstream of promoter), enrichment of active histone markers H3K4Me3 and H3K9Ac and subsequent POL2 recruitment, thus driving TERT transcription. Another study suggested a slightly different mechanism. According to work by Li et al.[85], the TERT promoter with C250T mutation was driven by NF-κB signaling. On activation of this signaling pathway, p52 (NF-κB2) is recruited to the C250T region, but not the C228T region, and cooperates with ETS factors ETS1/2 to drive efficient TERT transcription[85]. TERT promoter mutations are widely found together with BRAF V600E alteration in human cancers, particularly in thyroid cancer and melanoma[87-92]. A recent study found that that TERT promoter mutations and BRAF V600E cooperatively upregulated TERT expression and promoted the oncogenic behaviors in the papillary thyroid cancer cells[93].
Mechanisms of TERT promoter mutations in HCC
TERT promoter mutation was a later oncogenic event. Pilati et al.[94] have screened TERT promoter in a large series of liver cancers including adenomas, borderline lesions hepatocellular adenomas (HCA)/HCC, HCC derived from adenomas and classical HCC, and found TERT promoter mutations did not exist in classical adenomas, but in borderline lesions HCA/HCC (17%) and HCC cases derived from adenomas (56%) which frequency was similar to that in classical HCC (54%).
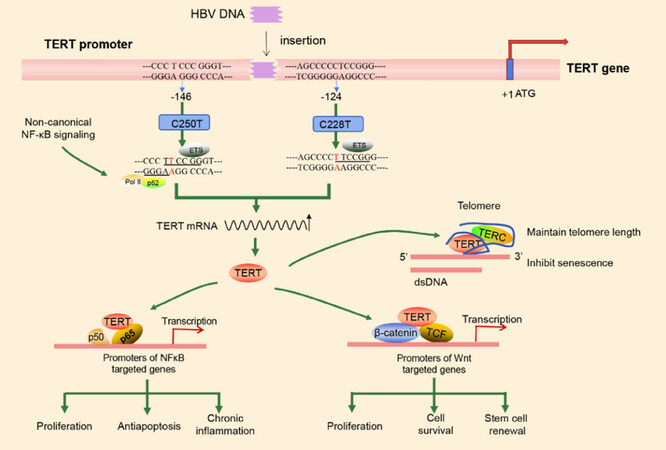
There are only a few studies focusing on the mechanism of how TERT promoter mutations influence TERT expression and lead to malignant transformation of liver cells [Figure 1]. Telomerase activation is important to maintaining telomere length that confers cancer cells infinite ability to overcome the proliferation barrier. One study demonstrated that TERT mRNA expression and telomerase activity were higher in patients with HCC who had both single nucleotide polymorphism (SNP) rs2853669 and promoter mutations of TERT gene[95]. The rs2853669 variant and the TERT promoter mutation C228T combined to induce TERT promoter methylation and increase TERT expression, resulting in a longer telomere length compared to the wild-type rs2853669 and TERT promoter[95].
Figure 1. Proposed model for telomerase reactivation by telomerase reverse transcriptase (TERT) promoter mutations. The C228T and C250T TERT promoter mutation both create an E-twenty-six (ETS) binding motif (the mutational hotspots are in red) to modulate TERT mRNA expression. P52 (NF-κB2) is recruited to the C250T region, but not the C228T region, and cooperates with ETS factors to drive efficient TERT transcription. The elevated TERT expression enhances cell malignant behavior through a telomere lengthening-dependent manner (maintaining telomere length or inhibiting senescence), and/or a telomere lengthening-independent manner (TERT acting as a transcriptional modulator regulating genes related to Wnt and NF-κB signaling pathways thereby promoting cell proliferation, antiapoptosis, and stem cell renewal). Hepatitis B virus (HBV) DNA insertion into TERT promoter is another possible mechanism of hepatocarcinogenesis, which may cause HBV promoter/enhancer-driven transcription of TERT
In recent years, TERT has been considered to have some other direct effects on carcinogenesis in addition to its function on maintaining telomere length[96]. Studies revealed that TERT acts as a transcriptional activator that activates the transcription of genes targeted by Wnt and NF-κB signaling to play a role in cell proliferation, antiapoptosis, and stem cell renewal[96,97]. In HCC, TERT expression level was higher in almost all cases with TERT promoter mutations than that in those without the mutations, and elevated TERT expression is closely related to the development of HCC[42,94]. Based on the significant association between TERT promoter and CTNNB1 mutations as well as previous studies showing the interaction between TERT and Wnt/β-catenin pathway, it was proposed that TERT promoter mutations and activation of the Wnt/β-catenin pathway together lead to malignant transformation[42,97]. By contrast, another research revealed that, while TERT expression did increase in the HCC cohort overall, it was not significantly correlated with TERT promoter mutations[48]. They suggested that TERT promoter mutations might cooperate with CDKN2A silencing to promote TERT mRNA expression. CDKN2A gene encodes the tumor suppressor gene p16INK4A, whose down-regulation together with up-regulated TERT expression is critical for epithelia cell immortalization[98]. Anyhow, there are only a few studies focusing on the mechanisms of TERT promoter mutations in HCC. Whether it shares the same mechanisms with other cancers requires further research in the future.
TERT promoter mutations in diagnosis, prognosis and therapy of HCC
A study detected the TERT promoter mutations in plasma cell-free DNA (cfDNA) in 218 patients with HCC, and the prevalence of TERT mutations was 47.7%, which was similar to the prevalence (44.4%) of 196 HCCs derived from the TCGA database[57]. Meanwhile, they also measured the prevalence of TERT promoter mutations in cfDNA of 81 patients with cirrhosis, and the frequency was 8.6%[57]. Since the frequency of TERT promoter mutations gradually increases during the process of cirrhosis and liver cancer, the TERT promoter mutations in the cfDNA in the serum can be detected as an important index for evaluating the development of HCC. However, there still remains a problem with specificity since the TERT promoter mutation is very common in various tumors so that the mutations in cfDNA cannot accurately reveal the source of the lesion.
The prognostic value of TERT promoter mutations remains controversial. Kawai-Kitahata et al.[7] and Huang et al.[43] performed survival analyses and demonstrated that TERT promoter mutations were associated with poor overall survival and could be prognostic markers for HCC[7,43]. However, Ko et al.[95] found that the presence of TERT promoter mutations alone did not translate into poor prognosis, but that the SNP rs2853669 and the -124C>T mutation combined were associated with poor survival rates. Further, Lee et al.[39] reported that longer telomere length, but not TERT promoter mutations, was independently associated with poor overall survival. Besides showing TERT promoter mutations’ correlation with poorer overall survival in HCC, Li et al.[99] also demonstrated that TERT amplifications were associated with shortened overall survival independent of other clinicopathological parameters such as age, gender and TNM staging. Thus, while we are sure that genetic changes at TERT gene have prognostic value, we are uncertain about exactly which factor(s) - TERT promoter mutations alone, the combination of the SNP rs2853669 and the -124C>T mutation, longer telomere length or TERT amplifications - directly indicate(s) poor prognosis.
It is believed that TERT is a promising but also challenging driver gene to target. There are no drugs specifically targeting TERT gene yet, although a few inhibitors have been used to target amplified genes in HCC: epidermal growth factor receptor inhibitors like Gefitinib targeting amplified EGFR, MET, MAPK1, MAPK3 and CRKL, Crizontinib and vemurafenib targeting BRAF and ERBB2, and alisertib targeting amplified AURKA[99]. According to Dhanasekaran et al.[100], the somatic mutations associated with liver tumor development lie in genes whose products are not easily or safely targeted, and that mutant TERT, TP53, CTNNB1, and MYC are even believed to be undruggable. Nevertheless, the study also reveals that a synthetic TERT DNA vaccine, INO-1400, is being tested in a phase 1 trial of patients with solid tumors (NCT02960594) and that some trials are using TERT promoter mutation as a biomarker for study enrollment (NCT02766270)[100]. Since a traditional strategy to target TERT is challenging, it is suggested that new strategies, such as microRNA-based therapeutics, should be developed to target driver genes like TERT or their pathways[100]. In fact, one study explored the potential of a novel immunotherapy using TERT-derived peptide (TERT461) as a vaccine by investigating its safety and immunogenicity and characterizing the TERT-specific T cell responses induced[101]. Their results showed that the vaccination induced TERT-specific immunity in 10/14 (71.4%) of the patients, and that 57.1% of patients treated with TERT461 peptide-specific T cells could prevent HCC recurrence after vaccination[101]. Another study also concluded that CypB, SART2, SART3, p53, MRP3, AFP, and TERT are promising tumor-associated antigens (TAAs) in HCC immunotherapy[102]. Besides, not only do they suggest that the administration of the TAAs or peptides containing their epitopes as vaccines after HCC treatment is likely to be effective, but they also demonstrated that the concurrent use of anti-CTLA-4 antibodies may further improve antitumor immunity[102]. Therefore, while it remains challenging to target TERT gene, new strategies are emerging to achieve this goal and make more effective therapy possible.
Conclusion
Our knowledge regarding the role of TERT promoter mutations in HCC is expanding; nevertheless, there remain many puzzles to be solved. Although the pattern of TERT promoter mutations in HCC is well-established, little is known about the mechanism through which TERT promoter mutations reactivate telomerase and promote tumor development. We are not yet sure how either somatic mutations or HBV integrations in the TERT promoter lead to malignant transformation and whether they can be prognostic biomarkers in HCC; nevertheless, we are confident that untangling the mechanisms relevant to TERT promoter can be a key for developing target therapy for HCC.
Declarations
Authors’ contributionsWriting the initial manuscript: Ma ZX, Yang CM
Revision of the manuscript: Yang CM, Ma ZX, Li MG
Drafting the outline of the manuscript, critical revision of the manuscript for intellectual content, finalizing the manuscript, and obtaining the funding: Tu H
Availability of data and materialsNot applicable.
Financial support and sponsorshipThis work was supported by grants from the National Natural Science Foundation of China (81572312); Chinese National Key Project Specialized for Infectious Diseases (2017ZX10201201-008-003); National Key Research and Development Program of China (2017YFC0908103); and Chinese State Key Laboratory of Oncogenes and Related Genes (91-14-16, 91-15-06).
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2019.
REFERENCES
1. Pezzuto F, Buonaguro L, Buonaguro FM, Tornesello ML. Frequency and geographic distribution of TERT promoter mutations in primary hepatocellular carcinoma. Infect Agent Cancer 2017;12:27.
2. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87-108.
3. Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med 2008;359:378-90.
5. Ringelhan M, Pfister D, O’Connor T, Pikarsky E, Heikenwalder M. The immunology of hepatocellular carcinoma. Nat Immunol 2018;19:222-32.
6. Zucman-Rossi J, Villanueva A, Nault JC, Llovet JM. Genetic landscape and biomarkers of hepatocellular carcinoma. Gastroenterology 2015;149:1226-39.e4.
7. Kawai-Kitahata F, Asahina Y, Tanaka S, Kakinuma S, Murakawa M, et al. Comprehensive analyses of mutations and hepatitis B virus integration in hepatocellular carcinoma with clinicopathological features. J Gastroenterol 2016;51:473-86.
8. Totoki Y, Tatsuno K, Covington KR, Ueda H, Creighton CJ, et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nat Genet 2014;46:1267-73.
9. Liu L, Lai S, Andrews LG, Tollefsbol TO. Genetic and epigenetic modulation of telomerase activity in development and disease. Gene 2004;340:1-10.
10. Lendvay TS, Morris DK, Sah J, Balasubramanian B, Lundblad V. Senescence mutants of Saccharomyces cerevisiae with a defect in telomere replication identify three additional EST genes. Genetics 1996;144:1399-412.
11. Lingner J, Cech TR. Purification of telomerase from Euplotes aediculatus: requirement of a primer 3’ overhang. Proc Natl Acad Sci U S A 1996;93:10712-7.
12. Cukusic A, Skrobot Vidacek N, Sopta M, Rubelj I. Telomerase regulation at the crossroads of cell fate. Cytogenet Genome Res 2008;122:263-72.
13. Pestana A, Vinagre J, Sobrinho-Simoes M, Soares P. TERT biology and function in cancer: beyond immortalisation. J Mol Endocrinol 2017;58:R129-R46.
14. Low KC, Tergaonkar V. Telomerase: central regulator of all of the hallmarks of cancer. Trends Biochem Sci 2013;38:426-34.
15. Liu T, Yuan X, Xu D. Cancer-specific telomerase reverse transcriptase (TERT) promoter mutations: biological and clinical implications. Genes (Basel) 2016;7:E38.
16. Daniel M, Peek GW, Tollefsbol TO. Regulation of the human catalytic subunit of telomerase (hTERT). Gene 2012;498:135-46.
17. Poole JC, Andrews LG, Tollefsbol TO. Activity, function, and gene regulation of the catalytic subunit of telomerase (hTERT). Gene 2001;269:1-12.
18. Greenberg RA, O’Hagan RC, Deng H, Xiao Q, Hann SR, et al. Telomerase reverse transcriptase gene is a direct target of c-Myc but is not functionally equivalent in cellular transformation. Oncogene 1999;18:1219-26.
19. Oh S, Song YH, Yim J, Kim TK. Identification of Mad as a repressor of the human telomerase (hTERT) gene. Oncogene 2000;19:1485-90.
20. Cerezo A, Kalthoff H, Schuermann M, Schafer B, Boukamp P. Dual regulation of telomerase activity through c-Myc-dependent inhibition and alternative splicing of hTERT. J Cell Sci 2002;115:1305-12.
21. Dwyer JM, Liu JP. Ets2 transcription factor, telomerase activity and breast cancer. Clin Exp Pharmacol Physiol 2010;37:83-7.
22. Wu KJ, Grandori C, Amacker M, Simon-Vermot N, Polack A, et al. Direct activation of TERT transcription by c-MYC. Nat Genet 1999;21:220-4.
23. Kanaya T, Kyo S, Hamada K, Takakura M, Kitagawa Y, et al. Adenoviral expression of p53 represses telomerase activity through down-regulation of human telomerase reverse transcriptase transcription. Clin Cancer Res 2000;6:1239-47.
24. Akincilar SC, Khattar E, Boon PL, Unal B, Fullwood MJ, et al. Long-range chromatin interactions drive mutant TERT promoter activation. Cancer Discov 2016;6:1276-91.
25. Barthel FP, Wei W, Tang M, Martinez-Ledesma E, Hu X, et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat Genet 2017;49:349-57.
26. Huang FW, Hodis E, Xu MJ, Kryukov GV, Chin L, et al. Highly recurrent TERT promoter mutations in human melanoma. Science 2013;339:957-9.
27. Alexiadis M, Rowley SM, Chu S, Leung DTH, Stewart CJR, et al. Mutational landscape of ovarian adult granulosa cell tumors from whole exome and targeted TERT promoter sequencing. Mol Cancer Res 2019;17:177-85.
28. Killela PJ, Reitman ZJ, Jiao Y, Bettegowda C, Agrawal N, et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci U S A 2013;110:6021-6.
29. Koelsche C, Sahm F, Capper D, Reuss D, Sturm D, et al. Distribution of TERT promoter mutations in pediatric and adult tumors of the nervous system. Acta Neuropathol 2013;126:907-15.
30. Allory Y, Beukers W, Sagrera A, Flandez M, Marques M, et al. Telomerase reverse transcriptase promoter mutations in bladder cancer: high frequency across stages, detection in urine, and lack of association with outcome. Eur Urol 2014;65:360-6.
31. Borah S, Xi L, Zaug AJ, Powell NM, Dancik GM, et al. Cancer. TERT promoter mutations and telomerase reactivation in urothelial cancer. Science 2015;347:1006-10.
32. Heidenreich B, Nagore E, Rachakonda PS, Garcia-Casado Z, Requena C, et al. Telomerase reverse transcriptase promoter mutations in primary cutaneous melanoma. Nat Commun 2014;5:3401.
33. Horn S, Figl A, Rachakonda PS, Fischer C, Sucker A, et al. TERT promoter mutations in familial and sporadic melanoma. Science 2013;339:959-61.
34. Hurst CD, Platt FM, Knowles MA. Comprehensive mutation analysis of the TERT promoter in bladder cancer and detection of mutations in voided urine. Eur Urol 2014;65:367-9.
35. Remke M, Ramaswamy V, Peacock J, Shih DJ, Koelsche C, et al. TERT promoter mutations are highly recurrent in SHH subgroup medulloblastoma. Acta Neuropathol 2013;126:917-29.
36. Vinagre J, Almeida A, Populo H, Batista R, Lyra J, et al. Frequency of TERT promoter mutations in human cancers. Nat Commun 2013;4:2185.
37. Wu S, Huang P, Li C, Huang Y, Li X, et al. Telomerase reverse transcriptase gene promoter mutations help discern the origin of urogenital tumors: a genomic and molecular study. Eur Urol 2014;65:274-7.
38. Campos MA, Macedo S, Fernandes M, Pestana A, Pardal J, et al. TERT promoter mutations are associated with poor prognosis in cutaneous squamous cell carcinoma. J Am Acad Dermatol 2019;80:660-9.e6.
39. Lee HW, Park TI, Jang SY, Park SY, Park WJ, et al. Clinicopathological characteristics of TERT promoter mutation and telomere length in hepatocellular carcinoma. Medicine (Baltimore) 2017;96:e5766.
40. Cevik D, Yildiz G, Ozturk M. Common telomerase reverse transcriptase promoter mutations in hepatocellular carcinomas from different geographical locations. World J Gastroenterol 2015;21:311-7.
41. Lee SE, Chang SH, Kim WY, Lim SD, Kim WS, et al. Frequent somatic TERT promoter mutations and CTNNB1 mutations in hepatocellular carcinoma. Oncotarget 2016;7:69267-75.
42. Nault JC, Mallet M, Pilati C, Calderaro J, Bioulac-Sage P, et al. High frequency of telomerase reverse-transcriptase promoter somatic mutations in hepatocellular carcinoma and preneoplastic lesions. Nat Commun 2013;4:2218.
43. Huang W, Zhou W, Li C, Yang Y, Shang YK, et al. Promoter mutations and cellular distribution of telomerase in non-clear cell and clear cell hepatocellular carcinoma. Oncotarget 2017;8:26288-97.
44. Yang XJ, Guo XC, Chen Y, Chen GR, Ma Y, et al. Telomerase reverse transcriptase promoter mutations in hepatitis B virus-associated hepatocellular carcinoma. Oncotarget 2016;7:27838-47.
45. Chen YL, Jeng YM, Chang CN, Lee HJ, Hsu HC, et al. TERT promoter mutation in resectable hepatocellular carcinomas: a strong association with hepatitis C infection and absence of hepatitis B infection. Int J Surg 2014;12:659-65.
46. Yuan XT, Cheng GH, Yu JY, Zheng SZ, Sun C, et al. The TERT promoter mutation incidence is modified by germline TERT rs2736098 and rs2736100 polymorphisms in hepatocellular carcinoma. Oncotarget 2017;8:23120-9.
47. Pezzuto F, Izzo F, Buonaguro L, Annunziata C, Tatangelo F, et al. Tumor specific mutations in TERT promoter and CTNNB1 gene in hepatitis B and hepatitis C related hepatocellular carcinoma. Oncotarget 2016;7:54253-62.
48. Cancer Genome Atlas Research Network. Electronic address: wheeler@bcm.edu; Cancer Genome Atlas Research Network. Comprehensive and Integrative Genomic Characterization of Hepatocellular Carcinoma. Cell 2017;169:1327-41.e23.
49. Nishida N, Nishimura T, Kaido T, Minaga K, Yamao K, et al. Molecular scoring of hepatocellular carcinoma for predicting metastatic recurrence and requirements of systemic chemotherapy. Cancers (Basel) 2018;10:E367.
50. Eichenmuller M, Trippel F, Kreuder M, Beck A, Schwarzmayr T, et al. The genomic landscape of hepatoblastoma and their progenies with HCC-like features. J Hepatol 2014;61:1312-20.
51. Fujimoto A, Furuta M, Totoki Y, Tsunoda T, Kato M, et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat Genet 2016;48:500.
52. Nault JC, Calderaro J, Di Tommaso L, Balabaud C, Zafrani ES, et al. Telomerase reverse transcriptase promoter mutation is an early somatic genetic alteration in the transformation of premalignant nodules in hepatocellular carcinoma on cirrhosis. Hepatology 2014;60:1983-92.
53. Calderaro J, Couchy G, Imbeaud S, Amaddeo G, Letouze E, et al. Histological subtypes of hepatocellular carcinoma are related to gene mutations and molecular tumour classification. J Hepatol 2017;67:727-38.
54. Donati B, Pietrelli A, Pingitore P, Dongiovanni P, Caddeo A, et al. Telomerase reverse transcriptase germline mutations and hepatocellular carcinoma in patients with nonalcoholic fatty liver disease. Cancer Med 2017;6:1930-40.
55. Dongiovanni P, Romeo S, Valenti L. Hepatocellular carcinoma in nonalcoholic fatty liver: role of environmental and genetic factors. World J Gastroenterol 2014;20:12945-55.
56. Ki Kim S, Ueda Y, Hatano E, Kakiuchi N, Takeda H, et al. TERT promoter mutations and chromosome 8p loss are characteristic of nonalcoholic fatty liver disease-related hepatocellular carcinoma. Int J Cancer 2016;139:2512-8.
57. Jiao J, Watt GP, Stevenson HL, Calderone TL, Fisher-Hoch SP, et al. Telomerase reverse transcriptase mutations in plasma DNA in patients with hepatocellular carcinoma or cirrhosis: prevalence and risk factors. Hepatol Commun 2018;2:718-31.
58. Jiang Z, Jhunjhunwala S, Liu J, Haverty PM, Kennemer MI, et al. The effects of hepatitis B virus integration into the genomes of hepatocellular carcinoma patients. Genome Res 2012;22:593-601.
59. Khoury JD, Tannir NM, Williams MD, Chen Y, Yao H, et al. Landscape of DNA virus associations across human malignant cancers: analysis of 3,775 cases using RNA-Seq. J Virol 2013;87:8916-26.
60. Paterlini-Brechot P, Saigo K, Murakami Y, Chami M, Gozuacik D, et al. Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene 2003;22:3911-6.
61. Tornesello ML, Buonaguro L, Izzo F, Buonaguro FM. Molecular alterations in hepatocellular carcinoma associated with hepatitis B and hepatitis C infections. Oncotarget 2016;7:25087-102.
62. Kaslow DC, Black S, Bloom DE, Datla M, Salisbury D, et al. Vaccine candidates for poor nations are going to waste. Nature 2018;564:337-9.
63. Yang X, Wu L, Lin J, Wang A, Wan X, et al. Distinct hepatitis B virus integration patterns in hepatocellular carcinoma and adjacent normal liver tissue. Int J Cancer 2017;140:1324-30.
64. Ferber MJ, Montoya DP, Yu C, Aderca I, McGee A, et al. Integrations of the hepatitis B virus (HBV) and human papillomavirus (HPV) into the human telomerase reverse transcriptase (hTERT) gene in liver and cervical cancers. Oncogene 2003;22:3813-20.
65. Sung WK, Zheng H, Li S, Chen R, Liu X, et al. Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat Genet 2012;44:765-9.
66. Gozuacik D, Murakami Y, Saigo K, Chami M, Mugnier C, et al. Identification of human cancer-related genes by naturally occurring hepatitis B virus DNA tagging. Oncogene 2001;20:6233-40.
67. Ding D, Lou X, Hua D, Yu W, Li L, et al. Recurrent targeted genes of hepatitis B virus in the liver cancer genomes identified by a next-generation sequencing-based approach. PLoS Genet 2012;8:e1003065.
68. Fujimoto A, Totoki Y, Abe T, Boroevich KA, Hosoda F, et al. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat Genet 2012;44:760-4.
69. Li W, Zeng X, Lee NP, Liu X, Chen S, et al. HIVID: an efficient method to detect HBV integration using low coverage sequencing. Genomics 2013;102:338-44.
70. Toh ST, Jin Y, Liu LZ, Wang JB, Babrzadeh F, et al. Deep sequencing of the hepatitis B virus in hepatocellular carcinoma patients reveals enriched integration events, structural alterations and sequence variations. Carcinogenesis 2013;34:787-98.
71. Lau CC, Sun T, Ching AK, He M, Li JW, et al. Viral-human chimeric transcript predisposes risk to liver cancer development and progression. Cancer Cell 2014;25:335-49.
72. Saitta C, Tripodi G, Barbera A, Bertuccio A, Smedile A, et al. Hepatitis B virus (HBV) DNA integration in patients with occult HBV infection and hepatocellular carcinoma. Liver Int 2015;35:2311-7.
73. Yan H, Yang Y, Zhang L, Tang G, Wang Y, et al. Characterization of the genotype and integration patterns of hepatitis B virus in early- and late-onset hepatocellular carcinoma. Hepatology 2015;61:1821-31.
74. Chiu YT, Wong JK, Choi SW, Sze KM, Ho DW, et al. Novel pre-mRNA splicing of intronically integrated HBV generates oncogenic chimera in hepatocellular carcinoma. J Hepatol 2016;64:1256-64.
75. Zhao LH, Liu X, Yan HX, Li WY, Zeng X, et al. Genomic and oncogenic preference of HBV integration in hepatocellular carcinoma. Nat Commun 2016;7:12992.
76. Lee WY, Bachtiar M, Choo CCS, Lee CG. Comprehensive review of hepatitis B virus-associated hepatocellular carcinoma research through text mining and big data analytics. Biol Rev Camb Philos Soc 2018; doi: 10.1111/brv.12457.
77. Schulze K, Imbeaud S, Letouze E, Alexandrov LB, Calderaro J, et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet 2015;47:505-11.
78. Iliopoulos D, Satra M, Drakaki A, Poultsides GA, Tsezou A. Epigenetic regulation of hTERT promoter in hepatocellular carcinomas. Int J Oncol 2009;34:391-9.
79. Kumakura S, Tsutsui TW, Yagisawa J, Barrett JC, Tsutsui T. Reversible conversion of immortal human cells from telomerase-positive to telomerase-negative cells. Cancer Res 2005;65:2778-86.
80. Widschwendter A, Muller HM, Hubalek MM, Wiedemair A, Fiegl H, et al. Methylation status and expression of human telomerase reverse transcriptase in ovarian and cervical cancer. Gynecol Oncol 2004;93:407-16.
81. Guilleret I, Benhattar J. Demethylation of the human telomerase catalytic subunit (hTERT) gene promoter reduced hTERT expression and telomerase activity and shortened telomeres. Exp Cell Res 2003;289:326-34.
82. Guilleret I, Yan P, Grange F, Braunschweig R, Bosman FT, et al. Hypermethylation of the human telomerase catalytic subunit (hTERT) gene correlates with telomerase activity. Int J Cancer 2002;101:335-41.
83. Dessain SK, Yu H, Reddel RR, Beijersbergen RL, Weinberg RA. Methylation of the human telomerase gene CpG island. Cancer Res 2000;60:537-41.
84. Zhang H, Weng X, Ye J, He L, Zhou D, et al. Promoter hypermethylation of TERT is associated with hepatocellular carcinoma in the Han Chinese population. Clin Res Hepatol Gastroenterol 2015;39:600-9.
85. Li Y, Zhou QL, Sun W, Chandrasekharan P, Cheng HS, et al. Non-canonical NF-kappaB signalling and ETS1/2 cooperatively drive C250T mutant TERT promoter activation. Nat Cell Biol 2015;17:1327-38.
86. Bell RJ, Rube HT, Kreig A, Mancini A, Fouse SD, et al. Cancer. The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 2015;348:1036-9.
87. Liu X, Bishop J, Shan Y, Pai S, Liu D, et al. Highly prevalent TERT promoter mutations in aggressive thyroid cancers. Endocr Relat Cancer 2013;20:603-10.
88. Xing M, Liu R, Liu X, Murugan AK, Zhu G, et al. BRAF V600E and TERT promoter mutations cooperatively identify the most aggressive papillary thyroid cancer with highest recurrence. J Clin Oncol 2014;32:2718-26.
89. Macerola E, Loggini B, Giannini R, Garavello G, Giordano M, et al. Coexistence of TERT promoter and BRAF mutations in cutaneous melanoma is associated with more clinicopathological features of aggressiveness. Virchows Arch 2015;467:177-84.
90. Liu R, Bishop J, Zhu G, Zhang T, Ladenson PW, et al. Mortality risk stratification by combining BRAF V600E and TERT promoter mutations in papillary thyroid cancer: genetic duet of BRAF and TERT promoter mutations in thyroid cancer mortality. JAMA Oncol 2016; doi: 10.1001/jamaoncol.2016.3288.
92. Nagore E, Heidenreich B, Rachakonda S, Garcia-Casado Z, Requena C, et al. TERT promoter mutations in melanoma survival. Int J Cancer 2016;139:75-84.
93. Liu R, Zhang T, Zhu G, Xing M. Regulation of mutant TERT by BRAF V600E/MAP kinase pathway through FOS/GABP in human cancer. Nat Commun 2018;9:579.
94. Pilati C, Letouze E, Nault JC, Imbeaud S, Boulai A, et al. Genomic profiling of hepatocellular adenomas reveals recurrent FRK-activating mutations and the mechanisms of malignant transformation. Cancer Cell 2014;25:428-41.
95. Ko E, Seo HW, Jung ES, Kim BH, Jung G. The TERT promoter SNP rs2853669 decreases E2F1 transcription factor binding and increases mortality and recurrence risks in liver cancer. Oncotarget 2016;7:684-99.
96. Ding D, Xi P, Zhou J, Wang M, Cong YS. Human telomerase reverse transcriptase regulates MMP expression independently of telomerase activity via NF-kappaB-dependent transcription. FASEB J 2013;27:4375-83.
97. Park JI, Venteicher AS, Hong JY, Choi J, Jun S, et al. Telomerase modulates Wnt signalling by association with target gene chromatin. Nature 2009;460:66-72.
98. Kiyono T, Foster SA, Koop JI, McDougall JK, Galloway DA, et al. Both Rb/p16INK4a inactivation and telomerase activity are required to immortalize human epithelial cells. Nature 1998;396:84-8.
99. Li X, Xu W, Kang W, Wong SH, Wang M, et al. Genomic analysis of liver cancer unveils novel driver genes and distinct prognostic features. Theranostics 2018;8:1740-51.
100. Dhanasekaran R, Nault JC, Roberts LR, Zucman-Rossi J. Genomic medicine and implications for hepatocellular carcinoma prevention and therapy. Gastroenterology 2019;156:492-509.
101. Mizukoshi E, Nakagawa H, Kitahara M, Yamashita T, Arai K, et al. Immunological features of T cells induced by human telomerase reverse transcriptase-derived peptides in patients with hepatocellular carcinoma. Cancer Lett 2015;364:98-105.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Ma ZX, Yang CM, Li MG, Tu H. Telomerase reverse transcriptase promoter mutations in hepatocellular carcinogenesis. Hepatoma Res 2019;5:8. http://dx.doi.org/10.20517/2394-5079.2018.104
AMA Style
Ma ZX, Yang CM, Li MG, Tu H. Telomerase reverse transcriptase promoter mutations in hepatocellular carcinogenesis. Hepatoma Research. 2019; 5: 8. http://dx.doi.org/10.20517/2394-5079.2018.104
Chicago/Turabian Style
Ma, Zi-Xian, Chun-Mei Yang, Meng-Ge Li, Hong Tu. 2019. "Telomerase reverse transcriptase promoter mutations in hepatocellular carcinogenesis" Hepatoma Research. 5: 8. http://dx.doi.org/10.20517/2394-5079.2018.104
ACS Style
Ma, Z.X.; Yang C.M.; Li M.G.; Tu H. Telomerase reverse transcriptase promoter mutations in hepatocellular carcinogenesis. Hepatoma. Res. 2019, 5, 8. http://dx.doi.org/10.20517/2394-5079.2018.104
About This Article
Special Issue
Copyright
Data & Comments
Data
 Cite This Article 45 clicks
Cite This Article 45 clicks












Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.