New strategy to distinguish clonal origin of RHCC/MHCC between intrahepatic metastasis and multicentric occurrence
Abstract
Hepatocellular carcinoma (HCC) is a kind of malignancy with high potential of metastasis and multicentric occurrence. The treatment of recurrent hepatocellular carcinoma (RHCC) and multinodular hepatocellular carcinoma (MHCC) is always a nodus because of the diverse clonal origin of RHCC/MHCC. Theoretically, the RHCC/MHCC can originate from intrahepatic metastasis (IM type) or multicentric occurrence (MO type). Our previous study proposed that there are at least 6 subtypes of clonal origin patterns in RHCC. RHCC and MHCC with different clonal origins have variant biological behaviors, clinical prognosis as well as treatment strategy. Generally speaking, patients with IM type HCC have a poorer prognosis compared with those with MO type HCC. Therefore, it is essential to emphasize the distribution of the clonal origin in HCC in order to determine the choice of clinical treatment. Undoubtedly, the detection of clonal origin pattern will become a promising breakthrough in the molecular pathological diagnosis of HCC. We should attach more attention to the establishment of a standardized molecular pathological clonal origin detection method and a new stratification of clinical treatment choice for RHCC/MHCC in future.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cancer related fatal diseases in the world, especially in China. The recent cancer statistics of China showed that its incidence was in the fourth place, and the mortality rate ranked the third[1]. With the development of time, the hepatic surgery has made great progress, and liver resection has become a routine method for the treatment of HCC[2]. However, the hepatic surgery is still facing two major obstacles. One is the treatment of recurrent hepatocellular carcinoma (RHCC). It was reported that the 5-year recurrence rate after hepatic resection of HCC is about 70% to 80%, or even higher[3-7]. Meanwhile, there is no consensus on the clinical treatment options for RHCC. Secondly, it is the treatment of multinodular hepatocellular carcinoma (MHCC). It has approved that the patient’s prognosis is poorer accompanied by the increased tumor nodules, especially > 3 foci[8]. One of the material causes for two major obstacles stems from the unprecise judgment of the clonal origin of RHCC and MHCC. It has affirmed the secondary tumor (synchronous or metachronous) was the core to directly reflect the biological behavior and determine patient’s prognosis[9-12]. For our practice, we found that two tumor nodules in one patient may have similar or different histological appearance, which may suggest the clonal origin of the tumors [Figure 1]. However, this judging method largely depends on the experience of the pathologist, which is not objective and accurate. Obviously, the clonal origin detection is unquestionably the check point to explore the biological behavior of HCC.
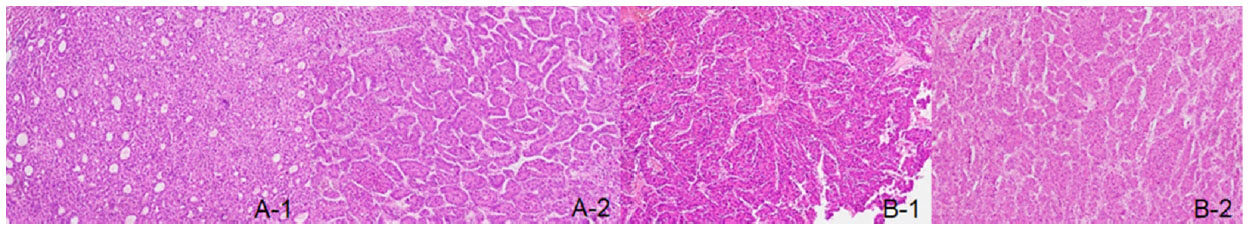
Figure 1. Hepatocellular carcinoma with different histological appearance and similar histological appearance. A-1: pseudoglandular pattern; A-2: thick trabecular pattern; B-1: thick trabecular pattern; B-2: thick trabecular pattern
HCC is a malignant tumor with high potential of recurrence and metastasis[13]. However, the clonal origin of RHCC/MHCC cannot be determined by simple clinical indicators and histopathology[14]. Consequently, the molecular pathological clonal origin detection is a new method to objectively determine the early, intermediate, and advanced stage of HCC in biological behavior and construct the basement of HCC molecular classification[15]. In other word, the clonal origin model directly affects the choice of clinical treatment.
Therefore, this review article briefly summarizes some relevant progresses of molecular pathological clonal origin of RHCC and MHCC. We searched all available publications regarding “clonal origin”, “recurrent hepatocellular carcinoma”, “multinodular hepatocellular carcinoma”, “intrahepatic metastasis”, and “multicentric occurrence” in the PubMed and focused the data mainly based on the high quality full-text format.
The clonal origin of HCC
The exploration of the clonal origin of the malignancy started in the blood system tumor[16,17]. Currently, it has approved that multiform clonal origins exist in malignant tumor. Identifying the clonal origin is of great significance for exploring tumor occurrence and evaluating tumor evolution[18-22]. For solidary tumor, there are two types of clonal origin, monoclonal origin and polyclonal origin[23]. Whether the secondary tumor is synchronous or metachronous, it may originate from intratumor metastasis of primary tumor (IM type); peradventure, it may be unrelated to the primary tumor, but from the normal cells which have adequate malignant mutation accumulation (MO type)[24]. Similarly, IM type HCC originates from the primary HCC with low degree of differentiation, incomplete envelope, widespread microvascular invasion (MVI) or even portal vein invasion. Among all of risk factors, MVI is considered to be the core factor in the occurrence of IM type HCC. According to our research on 686 HCC patients, the incidence of MVI was about 42%[25]. Remarkably, the incidence of MVI in single nodule HCC and MHCC are 40.4% and 55.6%, respectively. Higher incidence of MVI in MHCC indicates the possibility of IM type clonal origin in MHCC; MO type HCC is derived from the continuous blow of inflammation and fibrosis. Among the pathogenesis of inflammation, hepatitis viral is the most important reason and the most common cause of HCC. According to our statistics of 30 years’ HCC patients in the Eastern Hepatobiliary Surgery Hospital, the infection rate of hepatitis B virus (HBV) and hepatitis C virus (HCV) was 85.86% and 9.76%, respectively[26]. Therefore, effective inhibition of hepatitis virus replication is a key factor in the prevention of the occurrence of MO type HCC [Figure 2].
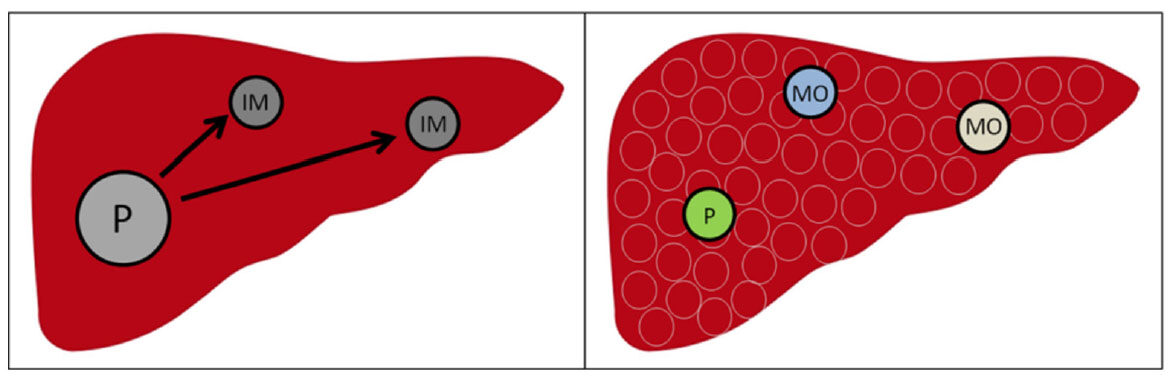
Figure 2. Mechanism of clonal origin with IM type and MO type in recurrent hepatocellular carcinoma/multinodular hepatocellular carcinoma. P: primary hepatocellular carcinoma; IM: intrahepatic metastasis; MO: multicentric occurrence
With the theory about the origination of malignant tumor constant improvement, such as tumor heterogeneity, cancer stem cells, circulating tumor cells, increased evidence suggests that there may be more complex clonal origin patterns in malignant tumor[27-29]. For example, heterogeneous clonal origin in single nodule HCC and IM-MO mixed clonal origin in RHCC and MHCC[30-32]. HCC with different clonal origin may engender variant clinical prognosis and therefore, different therapy method[33,34]. Consequently, it is a crucial cooperation for hepatic surgery and molecular pathology to formulate rational treatment strategy for RHCC and MHCC with different clonal origin.
The clonal origin of RHCC
The postoperative recurrence of HCC is likely to be an important indication of enhanced invasiveness of HCC and poor prognosis[35]. As a result, the current treatment strategy for primary HCC may not be suitable for RHCC. In view of this, scholars established many assessment systems for the prognosis of RHCC[36-41]. However, many studies focused on exploring the rational treatment of RHCC did not screen out the suitable groups for traditional treatments, such as hepatic resection, liver transplantation, transhepatic arterial chem otherapy and embolization (TACE), and radiofrequency ablation (RFA)[42-45]. It may attribute to the ignorance of great impact of clonal origin on the prognosis of patients.
Therefore, studies based on pathomorphology to predict the clonal origin of RHCC suggested that the incidence of IM type and MO type HCC is about 60% and 40%, respectively; IM type RHCC has poorer prognosis than MO type RHCC. Meanwhile, MO type RHCC and IM type RHCC are suitable for hepatic resection and TACE, respectively[46-48]. Based on above studies, to some extent, it is meaningful to judge the clonal origin of RHCC by histopathology. However, the experience of pathologist may affect the judgment of the clonal origin pattern. Therefore, histopathology cannot objectively and quantitatively reflect the real biological behavior of RHCC. To sum up, it is necessary for us to establish therapeutic strategy for RHCC with different clonal origin according to molecular pathological examination, so as to enable patients to get the best prognosis.
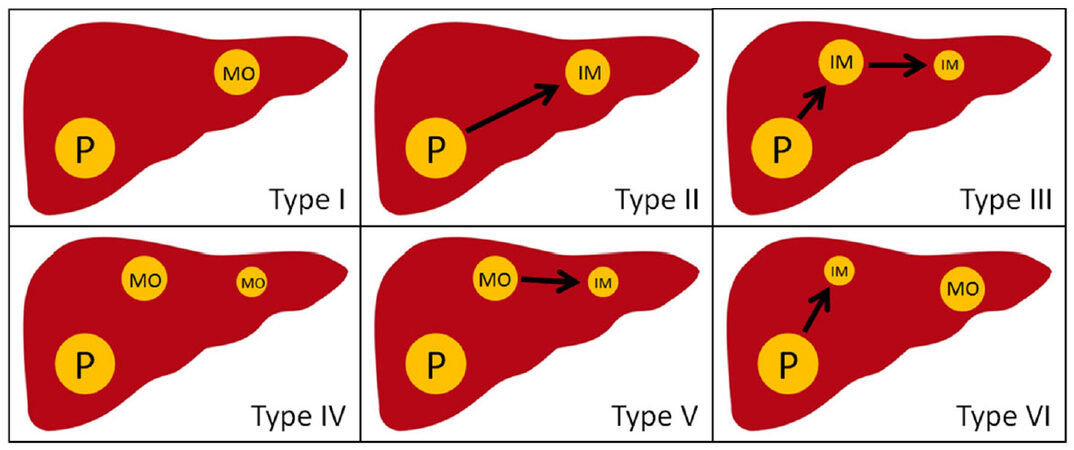
Molecular pathology applies a variety of methods to determine the clonal origin of RHCC. The HBV infection is present in most patients with HCC. Chen et al.[49] used southern-blot to detect the hepatitis B virus DNA (HBV-DNA) integration site in 5 cases of RHCC. Compared with 2 cases of IM type, 3 cases were MO type. Yamamoto et al.[50] checked the HBV-DNA integration site and its flanking genomic DNA, and found that 6 of 8 cases of RHCC were MO type and 2 were IM type. Interestingly, Liang et al.[51] used the same method, and found that, for multiple nodular RHCC, there are some nodules with the same clonal origin of primary HCC while other nodules is different, which is IM-MO mixed type RHCC. These studies provide a basis for the study of the clonal origin pattern of RHCC. However, HBV-DNA integration site detection is only suitable for HBV-related HCC. Referring to the distributed gene expression between primary HCC and RHCC, the scholars explored the clonal origin of RHCC by DNA ploidy analysis and p53 gene mutation site analysis[52-54]. However, the case of RHCC in these studies is little (< 20 patients). Moreover, the above studies only explained two clonal origin patterns of RHCC, but not integrated with prognosis of patients. Therefore, we adopted microdissecton-based PCR single-strand conformation polymorphism assay to check fifteen high-frequency of loss of heterozygosity (LOH) of DNA microsatellites on 100 tumor nodules in 60 matched pairs of RHCC from 40 patients who underwent liver re-resection. The definitions of the MO type and the IM type of RHCC were as follows: a ≥ 30% difference (number of different LOH loci/number of informative loci × 100) between primary HCC and any recurrent nodule was defined as MO type, on the contrary, IM type. Among all the patients, the percentage of IM type RHCC and MO type RHCC was 76.7% and 23.3%, respectively. MO type RHCC had a better prognosis than IM type RHCC (OS 130.8 ± 8.5 months vs. 80.8 ± 8.5 months; RFS 33.8 ± 4.5 months vs. 14.2 ± 2.5 months)[33]. Then, we classified 2 clonal patterns into 6 subclonal types: type I, single-nodular MO-RHCC; type II, single-nodular IM-RHCC; type III, single-nodular IM-RHCC spreading intrahepatic metastasis; type IV, multinodular MO-RHCC; type V, single-nodular MO-RHCC spreading intrahepatic metastasis; and type VI, single-nodular MO-RHCC combined with IM-RHCC [Figure 3]. Among them, type I, IV, and VI is MO type; Type II, III, and V is IM type. We recommended liver re-resection for MO type RHCC, and interventional therapy for IM type RHCC. This classification provided a theoretical basis for the selection of clinical treatment.
Figure 3. Six subtypes of clonal origin in recurrent hepatocellular carcinoma/multinodular hepatocellular carcinoma. P: primary hepatocellular carcinoma; IM: intrahepatic metastasis; MO: multicentric occurrence
With the development of the next-generation sequencing technology, we can explore the clonal origin of RHCC from the level of the whole genome expression spectrum. Shi et al.[55] sequenced the whole exome with 1 case of RHCC patient of MHCC after resection. The gene expression profile of two RHCC nodules was highly similar with one primary nodule (86.7% and 86.6% respectively), rather than other primary nodule which pointed out the clonal origin of RHCC.
The clonal origin of MHCC
MHCC is a common clinical form of HCC. At present, scholars in various countries, including some international standards, have not yet reached a consensus on the clinical diagnosis and staging of MHCC. For example, there is controversy about ≥ 2 nodules or ≥ 3 nodules as the standard of MHCC[56]. The Barcelona clinic liver cancer (BCLC) staging classification defined ≤ 3 nodules, ≤ 3 cm as stage A, called the early stage; ≥ 4 tumors of any size, or > 3 cm, 2-3 tumors are classified as stage B, called the intermediate stage, and defined as MHCC[57]. Therefore, MHCC is not considered as early form of HCC in BCLC staging classification. Accordingly, the guidelines of HCC in Europe and America also recommend TACE/sorafenib as a first-line treatment for MHCC[58,59]. However, if such kind of HCC occurred based on clonal origin of MO type, then they should not be considered pathobiologically as in the intermediate progression stage, and their treatment strategy will also be different accordingly. As the exploration of different treatment with BCLC intermediate stage of HCC, hepatic resection for some patients can obtain better prognosis than conservative treatment[60,61].
With the increase of nodule and the scattered nodule, the prognosis of the patients is worse[62-64]. Therefore, the current clinical study is paying more attention to the screening of radical treatment for MHCC[65,66]. Huang et al.[67] studied 102 MHCC patients with less than 3 nodules, and found that the presence of MVI is an independent risk factor for the patients of early recurrence (< 1 year) (HR, 4.02, 95% CI, 1.42-11.39, P = 0.009). Nojiri et al.[68] retrospectively analyzed 107 patients of MHCC who underwent R0 resection and found that, for the patients with > 4 nodules, vascular invasion was an independent risk factor for long-term survival (1-year overall survival 71.1% vs. 82.4%, 3-year overall survival 36.9% vs. 61%, 5-year overall survival 0% vs. 25.4%, P = 0.0035). In view of vascular invasion, it is an important indication for the occurrence of MHCC as IM type. To sum up, no matter the number of nodules, vascular invasion are always the important prognostic factors for MHCC. Referring to the correlation between vascular invasion and IM type clonal origin, effective screening of MO type MHCC patients for actively radical treatment has become an important point of MHCC clonal origin research.
Similar to the research of RHCC clonal origin, the study of MHCC clonal origin also begins with the HBV-DNA integration site analysis. Govindarajan et al.[69] and Aoki et al.[70] analyzed the HBV-DNA integration sites in 2 cases of MHCC, respectively, and preliminarily established the concept of IM type and MO type in MHCC. After that, some scholars used different methods, such as analysis of methylation pattern of X-chromosome-linked human androgen receptor gene, mitochondrial D-loop mutations analysis, DNA fingerprinting analysis, analysis of difference of tumor suppressor gene promoter region methylation, to confirm the existence of IM type and MO type MHCC[71-74]. Subsequently, scholars began to pay attention to the proportion of IM type and MO type in MHCC. Hsu et al.[75] analyzed the HBV-DNA integration site of 25 cases of MHCC, including the main tumor, satellites and metastatic loci, and found that the IM type and MO type accounted for 60.7% and 39.3%, respectively. Tsuda et al.[76] detected the alleles LOH of chromosome 16 in 19 MHCC patients, and found that the IM type and MO type accounted for 52.4% and 47.6%, respectively. Hui et al.[77] performed DNA ploidy analysis of 62 tumor nodules in 26 MHCC patients, and found that IM type and MO type accounted for 53.8% and 46.2%, respectively. Based on our detection of the clonal origin of 439 cases of MHCC in Eastern Hepatobiliary Surgery Hospital, IM type and MO type MHCC account for 51.9% and 48.1%, respectively (unpublished data). Referring to the clonal origin of RHCC, we believe that MHCC is likely to have the same clonal origin patterns with RHCC [Figure 3]. Therefore, the choices of clinical treatment patterns for patients with MHCC should be based on the clonal origin patterns of MHCC in order to get better prognosis for these patients.
With the development of the next-generation sequencing technology, the understanding of clonal origin of MHCC can be penetrated into the level of specific gene and whole gene expression profiles. Xue et al.[31] performed exome and low-depth, whole-genome sequencing for 43 nodules of primary tumors, satellite foci, metastatic foci and multiple foci in 10 patients with MHCC. They found that the proportion of ubiquitous mutations in different tumor nodules in the same patient varied with 8%-97%. Furuta et al.[78] performed whole genome sequencing and RNA sequencing for 49 nodules from 23 MHCC patients, which provides more detailed genetic information for clonal origin of MHCC. Lin et al.[79] applied the whole exome sequencing to analyse 69 lesions from 11 MHCC patients, and found that 29% of driver mutations is heterogeneous. The heterogeneity of methylation level may be a key for the occurrence and progress of MHCC.
Techniques of clonal origin detection
The criteria for judging the clonal origin of IM type and MO type HCC have not been widely accepted. Some studies based on whether the recurrent time < 1 year or histopathology to define IM type and MO type RHCC[6,47]. However, these classification methods can not accurately and objectively reflect the clonal origin of HCC. Therefore, molecular pathology uses a variety of methods to confirm it: HBV-DNA integration site analysis, DNA ploidy analysis, DNA fingerprint analysis, X-chromosome inactivation pattern detection, chromosomal LOH analysis, p53 gene mutation analysis, mitochondrial D-loop mutations analysis, microsatellite LOH analysis, next-generation sequencing technology, and so on [Table 1]. Some scholars has compared various kinds of methods[80,81]. According to our experience, we recommended the microsatellite LOH detection[33,82,83]. It is not only suitable for paraffin embedded tissues, resolves the restriction of gender and HBV infection, but also it can select a set of microsatellite profile to improve the diagnostic accuracy. In addition, microsatellite DNA is a suitable marker to reflect the overall stability of genome. To sum up, microsatellite LOH detection is the relatively ideal method to reduce the bias of HCC heterogeneity to the clonal origin in various methods.
Techniques of clonal origin detection
| Technique | Method | Material | Genomic loci | Reference |
|---|---|---|---|---|
| HBV-DNA integration pattern | Southern blot analysis | Freshly frozen tissue | HBV DNA | [49,51,69,70,75] |
| HBV-DNA and flanking human DNA junctions | PCR | Paraffin-embedded tissue | HBV DNA | [50] |
| DNA fingerprint analysis | AP-PCR | Paraffin-embedded tissue | Nuclear DNA | [72] |
| DNA ploidy analysis | Feulgen-DNA analysis; flow cytometric method | Paraffin-embedded tissue | Nuclear DNA | [52,53,77] |
| X-chromosome inactivation pattern | PCR | Freshly frozen tissue | The HUMARA locus of exon 1 of the X-chromosomelinked human androgen receptor gene | [74] |
| Chromosomal alterations | Comparative genomic hybridization | Freshly frozen tissue | Nuclear chromosome | [80] |
| Chromosomal LOH | RFLP analysis | Freshly frozen tissue | HBA1, D16S32, D16S34, D16S35, CETP, MT2, D16S4, HP, TAT, CTRB, APRT | [76] |
| Mitochondrial D-loop mutations | PCR | Freshly frozen tissue | Mitochondrial DNA D-loop region | [73] |
| Allelotype and LOH of p53 gene | BanII RFLP analysis | Freshly frozen tissue | Sequencing of exons 5, 7, and 8 of the TP53 gene | [54] |
| Microsatellite LOH | PCR | Paraffin-embedded tissue | D1S243, D1S507, D4S402, D4D406, D4S415, D8S264, D8S277, D8S520, D13S268, et al. | [33,34,82,83] |
| Tumor genomic heterogeneity analysis | Next-generation sequencing technology | Freshly frozen tissue | Whole-genome sequencing | [31,55,78,79] |
Conclusion
With the development of time, the molecular biological behavior and characteristics of HCC has become an important guide for hepatic surgery. Among them, RHCC and MHCC will be an important breakthrough in improving the long-term effect of HCC. Molecular cloning detection is an important theoretical and technical support to break this bottleneck. Therefore, strengthening the study of clonal origin of HCC and establishing a scientific and precise molecular cloning detection technology will be an important task in the field of HCC pathology. The innovation of molecular cloning technology provides guidance for the individualized treatment strategy of RHCC and MHCC. The overall view is that the IM type HCC has a more malignant biological behavior, and poorer clinical prognosis than the MO type HCC, no matter RHCC or MHCC.
The molecular pathological technical standards for evaluating the clonal origin of HCC have not yet been unified. Microsatellite LOH detection is currently the most widely used method in clinical practice. We should explore the method to unite high sensitivity and specificity, low cost, convenient and quick to serve the clinical practice better in future.
Declarations
Authors’ contributionsReviewed the literature and wrote the manuscript: Wang H, Cong WM
Financial support and sponsorshipNone.
Conflicts of interestThere are no conflicts of interest.
Patient consentNot applicable.
Ethics approvalNot applicable.
Copyright© The Author(s) 2018.
REFERENCES
1. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. Cancer J Clin 2016;66:115-32.
2. Bruix J, Gores GJ, Mazzaferro V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut 2014;63:844-55.
3. Belghiti J, Panis Y, Farges O, Benhamou JP, Fekete F. Intrahepatic recurrence after resection of hepatocellular carcinoma complicating cirrhosis. Ann Surg 1991;214:114-7.
4. Imamura H, Matsuyama Y, Miyagawa Y, Ishida K, Shimada R, Miyagawa S, Makuuchi M, Kawasaki S. Prognostic significance of anatomical resection and des-gamma-carboxy prothrombin in patients with hepatocellular carcinoma. Br J Surg 1999;86:1032-8.
5. Grazi GL, Ercolani G, Pierangeli F, Del Gaudio M, Cescon M, Cavallari A, Mazziotti A. Improved results of liver resection for hepatocellular carcinoma on cirrhosis give the procedure added value. Ann Surg 2001;234:71-8.
6. Poon RT, Fan ST, Ng IO, Lo CM, Liu CL, Wong J. Different risk factors and prognosis for early and late intrahepatic recurrence after resection of hepatocellular carcinoma. Cancer 2000;89:500-7.
7. Mise Y, Hasegawa K, Shindoh J, Ishizawa T, Aoki T, Sakamoto Y, Sugawara Y, Makuuchi M, Kokudo N. The feasibility of third or more repeat hepatectomy for recurrent hepatocellular carcinoma. Ann Surg 2015;262:347-57.
8. Giannini EG, Farinati F, Ciccarese F, Pecorelli A, Rapaccini GL, Di Marco M, Benvegnu L, Caturelli E, Zoli M, Borzio F, Chiaramonte M, Trevisani F; Italian Liver Cancer Group. Prognosis of untreated hepatocellular carcinoma. Hepatology 2015;61:184-90.
9. Liu G, Wang K, Li J, Xia Y, Lu L, Wan X, Yan Z, Shi L, Lau WY, Wu M, Shen F. Changes in serum alpha fetoprotein in patients with recurrent hepatocellular carcinoma following hepatectomy. J Gastroenterol Hepatol 2015;30:1405-11.
10. Hou YF, Wei YG, Yang JY, Wen TF, Xu MQ, Yan LN, Li B, Chen KF. Microvascular invasion patterns affect survival in hepatocellular carcinoma patients after second hepatectomy. J Surg Res 2016;200:82-90.
11. Chen R, Gan Y, Ge N, Chen Y, Wang Y, Zhang B, Wang Y, Ye S, Ren Z. Transarterial chemoembolization versus radiofrequency ablation for recurrent hepatocellular carcinoma after resection within Barcelona clinic liver cancer stage 0/A: a retrospective comparative study. J Vasc Interv Radiol 2016;27:1829-36.
12. Sun WC, Chen IS, Liang HL, Tsai CC, Chen YC, Wang BW, Lin HS, Chan HH, Hsu PI, Tsai WL, Cheng JS. Comparison of repeated surgical resection and radiofrequency ablation for small recurrent hepatocellular carcinoma after primary resection. Oncotarget 2017;8:104571-81.
14. Cong WM, Bu H, Chen J, Dong H, Zhu YY, Feng LH, Chen J. Practice guidelines for the pathological diagnosis of primary liver cancer: 2015 update. World J Gastroenterol 2016;22:9279-87.
15. Cong WM, Wu MC. New insights into molecular diagnostic pathology of primary liver cancer: advances and challenges. Cancer letters 2015;368:14-9.
16. Fialkow PJ, Gartler SM, Yoshida A. Clonal origin of chronic myelocytic leukemia in man. Proc Natl Acad Sci 1967;58:1468-71.
17. Fialkow PJ, Klein G, Gartler SM, Clifford P. Clonal origin for individual Burkitt tumours. Lancet 1970;1:384-6.
18. Van Etten JL, Dehm SM. Clonal origin and spread of metastatic prostate cancer. Endocr Relat Cancer 2016;23:R207-17.
19. Begg CB, Ostrovnaya I, Carniello JV, Sakr RA, Giri D, Towers R, Schizas M, De Brot M, Andrade VP, Mauguen A, Seshan VE, King TA. Clonal relationships between lobular carcinoma in situ and other breast malignancies. Breast Cancer Res 2016;18:66.
20. Baker AM, Graham TA, Wright NA. Pre-tumour clones, periodic selection and clonal interference in the origin and progression of gastrointestinal cancer: potential for biomarker development. J Pathol 2013;229:502-14.
21. Castellarin M, Milne K, Zeng T, Tse K, Mayo M, Zhao Y, Webb JR, Watson PH, Nelson BH, Holt RA. Clonal evolution of high-grade serous ovarian carcinoma from primary to recurrent disease. J Pathol 2013;229:515-24.
22. Shin K, Lim A, Odegaard JI, Honeycutt JD, Kawano S, Hsieh MH, Beachy PA. Cellular origin of bladder neoplasia and tissue dynamics of its progression to invasive carcinoma. Nature Cell Biol 2014;16:469-78.
24. Nomoto S, Hishida M, Inokawa Y, Sugimoto H, Kodera Y. Management of hepatocellular carcinoma should consider both tumor factors and background liver factors. Hepatobil Surg Nutr 2014;3:82-5.
25. Feng LH, Dong H, Lau WY, Yu H, Zhu YY, Zhao Y, Lin YX, Chen J, Wu MC, Cong WM. Novel microvascular invasion-based prognostic nomograms to predict survival outcomes in patients after R0 resection for hepatocellular carcinoma. J Cancer Res Clin Oncol 2017;143:293-303.
26. Cong WM, Dong H, Zhu YY, Zhu Z. Malignant tumors of the liver and intrahepatic bile ducts. In: Cong WM, editor. Surgical pathology of hepatobiliary tumors Singapore: Springer Nature Singapore; 2017. pp. 145-8.
27. Massague J, Obenauf AC. Metastatic colonization by circulating tumour cells. Nature 2016;529:298-306.
28. McGranahan N, Swanton C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 2017;168:613-28.
29. Lasorella A, Benezra R, Iavarone A. The ID proteins: master regulators of cancer stem cells and tumour aggressiveness. Nature Rev Cancer 2014;14:77-91.
30. Moeini A, Sia D, Zhang Z, Camprecios G, Stueck A, Dong H, Montal R, Torrens L, Martinez-Quetglas I, Fiel MI, Hao K, Villanueva A, Thung SN, Schwartz ME, Llovet JM. Mixed hepatocellular cholangiocarcinoma tumors: cholangiolocellular carcinoma is a distinct molecular entity. J Hepatol 2017;66:952-61.
31. Xue R, Li R, Guo H, Guo L, Su Z, Ni X, Qi L, Zhang T, Li Q, Zhang Z, Xie XS, Bai F, Zhang N. Variable intra-tumor genomic heterogeneity of multiple lesions in patients with hepatocellular carcinoma. Gastroenterology 2016;150:998-1008.
32. Duan M, Hao J, Cui S, Worthley DL, Zhang S, Wang Z, Shi J, Liu L, Wang X, Ke A, Cao Y, Xi R, Zhang X, Zhou J, Fan J, Li C, Gao Q. Diverse modes of clonal evolution in HBV-related hepatocellular carcinoma revealed by single-cell genome sequencing. Cell Res 2018;28:359-73.
33. Wang B, Xia CY, Lau WY, Lu XY, Dong H, Yu WL, Jin GZ, Cong WM, Wu MC. Determination of clonal origin of recurrent hepatocellular carcinoma for personalized therapy and outcomes evaluation: a new strategy for hepatic surgery. J Am Coll Surg 2013;217:1054-62.
34. Wang Z, Gong W, Shou D, Zhang L, Gu X, Wang Y, Teng D, Zheng H. Clonal origin of hepatocellular carcinoma and recurrence after liver transplantation. Ann Transplant 2016;21:484-90.
35. de'Angelis N, Landi F, Carra MC, Azoulay D. Managements of recurrent hepatocellular carcinoma after liver transplantation: a systematic review. World J Gastroenterol 2015;21:11185-98.
36. Choi SI, Yu A, Kim BH, Ko EJ, Park SS, Nam BH, Park JW. A model predicting survival of patients with recurrent or progressive hepatocellular carcinoma: the MORE score. J Gastroenterology Hepatol 2017;32:651-8.
37. Bodzin AS, Lunsford KE, Markovic D, Harlander-Locke MP, Busuttil RW, Agopian VG. Predicting mortality in patients developing recurrent hepatocellular carcinoma after liver transplantation: impact of treatment modality and recurrence characteristics. Ann Surg 2017;266:118-25.
38. Nagai S, Mangus RS, Kubal CA, Ekser B, Fridell JA, Klingler KR, Maluccio MA, Tector AJ. Prognosis after recurrence of hepatocellular carcinoma in liver transplantation: predictors for successful treatment and survival. Clin Transplant 2015;29:1156-63.
39. Na GH, Hong TH, You YK, Kim DG. Clinical analysis of patients with hepatocellular carcinoma recurrence after living-donor liver transplantation. World J Gastroenterol 2016;22:5790-9.
40. Muaddi H, Al-Adra DP, Beecroft R, Ghanekar A, Moulton CA, Doyle A, Selzner M, Wei A, McGilvray ID, Gallinger S, Grant DR, Cattral MS, Greig PD, Kachura J, Cleary SP, Sapisochin G. Liver transplantation is equally effective as a salvage therapy for patients with hepatocellular carcinoma recurrence following radiofrequency ablation or liver resection with curative intent. Ann Surg Oncol 2018;25:991-9.
41. Perea Del Pozo E, Bernal Bellido C, Sendin Matin M, Cepeda Franco C, Alamo Martinez JM, Suarez Artacho G, Marin Gomez LM, Padillo Ruiz J, Gomez Bravo MA. Recurrent hepatocellular carcinoma after liver transplantation: analysis of risk factors. Transplant Proc 2016;48:2990-3.
42. Erridge S, Pucher PH. Meta-analysis of determinants of survival following treatment of recurrent hepatocellular carcinoma. Br J Surg 2017;104:1433-42.
43. Gavriilidis P, Askari A, Azoulay D. Survival following redo hepatectomy vs radiofrequency ablation for recurrent hepatocellular carcinoma: a systematic review and meta-analysis. HPB (Oxford) 2017;19:3-9.
44. Wang DY, Liu L, Qi XS, Su CP, Chen X, Liu X, Chen J, Li HY, Guo XZ. Hepatic re-resection versus transarterial chemoembolization for the treatment of recurrent hepatocellular carcinoma after initial resection: a systematic review and meta-analysis. Asian Pac J Cancer Prev 2015;16:5573-8.
45. Huang J, Yan L, Wu H, Yang J, Liao M, Zeng Y. Is radiofrequency ablation applicable for recurrent hepatocellular carcinoma after liver transplantation? J Surg Res 2016;200:122-30.
46. Zhang X, Li C, Wen T. Outcomes of salvage liver transplantation and re-resection/radiofrequency ablation for intrahepatic recurrent hepatocellular carcinoma: a new surgical strategy based on recurrence pattern. Dig Dis Sci 2018;63:502-14.
47. Hao S, Fan P, Chen S, Tu C, Wan C. Distinct recurrence risk factors for intrahepatic metastasis and multicenter occurrence after surgery in patients with hepatocellular carcinoma. J Gastrointest Surg 2017;21:312-20.
48. Zhang X, Li C, Wen T, Yan L, Li B, Yang J, Wang W, Xu M, Lu W, Jiang L. Appropriate treatment strategies for intrahepatic recurrence after curative resection of hepatocellular carcinoma initially within the Milan criteria: according to the recurrence pattern. Eur J Gastroenterol Hepatol 2015;27:933-40.
49. Chen PJ, Chen DS, Lai MY, Chang MH, Huang GT, Yang PM, Sheu JC, Lee SC, Hsu HC, Sung JL. Clonal origin of recurrent hepatocellular carcinomas. Gastroenterology 1989;96:527-9.
50. Yamamoto T, Kajino K, Kudo M, Sasaki Y, Arakawa Y, Hino O. Determination of the clonal origin of multiple human hepatocellular carcinomas by cloning and polymerase chain reaction of the integrated hepatitis B virus DNA. Hepatology 1999;29:1446-52.
51. Liang XH. Clonal origin of intrahepatic recurrence after resection of hepatocellular carcinoma. Chin J Oncol 1991;13:2-4.
52. Kuo SH, Sheu JC, Chen DS, Sung JL, Lin CC, Hsu HC. DNA clonal heterogeneity of hepatocellular carcinoma demonstrated by Feulgen-DNA analysis. Liver 1987;7:359-63.
53. Nagasue N, Kohno H, Chang YC, Yamanoi A, Kimoto T, Takemoto Y, Nakamura T. DNA ploidy pattern in synchronous and metachronous hepatocellular carcinomas. J Hepatol 1992;16:208-14.
54. Hsu HC, Peng SY, Lai PL, Sheu JC, Chen DS, Lin LI, Slagle BL, Butel JS. Allelotype and loss of heterozygosity of p53 in primary and recurrent hepatocellular carcinomas. A study of 150 patients. Cancer 1994;73:42-7.
55. Shi JY, Xing Q, Duan M, Wang ZC, Yang LX, Zhao YJ, Wang XY, Liu Y, Deng M, Ding ZB, Ke AW, Zhou J, Fan J, Cao Y, Wang J, Xi R, Gao Q. Inferring the progression of multifocal liver cancer from spatial and temporal genomic heterogeneity. Oncotarget 2016;7:2867-77.
56. Cong WM. The status and thinking of clinicopathological study of multiple nodular hepatocellular carcinoma. Available from: http://oversea.cnki.net/kcms/detail/detail.aspxrecid=&FileName=ZGKA201510001011&DbName=CPFD2016&DbCode=CPFD [Last accessed on 24 May 2018].
57. Llovet JM, Brú C, Bruix J. Prognosis of hepatocellular carcinoma: the BCLC staging classification. Semin Liver Dis 1999;19:329-38.
58. European Association for the Study of the Liver. EASL-EORTC clinical practice guidelines: management of hepatocellular carcinoma. J Hepatol 2012;56:908-43.
59. Bruix J, Sherman M. Management of hepatocellular carcinoma: an update. Hepatology 2011;53:1020-2.
60. Wada H, Eguchi H, Noda T, Ogawa H, Yamada D, Tomimaru Y, Tomokuni A, Asaoka T, Kawamoto K, Gotoh K, Marubashi S, Umeshita K, Nagano H, Doki Y, Mori M. Selection criteria for hepatic resection in intermediate-stage (BCLC stage B) multiple hepatocellular carcinoma. Surgery 2016;160:1227-35.
61. Zhong JH, Ke Y, Gong WF, Xiang BD, Ma L, Ye XP, Peng T, Xie GS, Li LQ. Hepatic resection associated with good survival for selected patients with intermediate and advanced-stage hepatocellular carcinoma. Ann Surg 2014;260:329-40.
62. Torzilli G, Belghiti J, Kokudo N, Takayama T, Capussotti L, Nuzzo G, Vauthey JN, Choti MA, De Santibanes E, Donadon M, Morenghi E, Makuuchi M. A snapshot of the effective indications and results of surgery for hepatocellular carcinoma in tertiary referral centers: is it adherent to the EASL/AASLD recommendations? An observational study of the HCC east-west study group. Ann Surg 2013;257:929-37.
63. Qin HG, Zhong JH, Xiang BD, Wu FX, Peng NF, You XM, Yuan WP, Gong WF, Ma L, Li LQ. Comparison of the prognosis after hepatic resection for patients with Barcelona Clinical Liver Cancer stage B hepatocellular carcinoma. Natl Med J China 2016;96:3384-8.
64. Zhou YD, Li HK, Cui YL, Zhang T, Li Q. The indications for hepatectomy for multinodular hepatocellular carcinoma: experience from a single institution. Dig Surg 2015;32:82-9.
65. Ciria R, Lopez-Cillero P, Gallardo AB, Cabrera J, Pleguezuelo M, Ayllon MD, Luque A, Zurera L, Espejo JJ, Rodriguez-Peralvarez M, Montero JL, de la Mata M, Brice-o J. Optimizing the management of patients with BCLC stage-B hepatocellular carcinoma: modern surgical resection as a feasible alternative to transarterial chemoemolization. Eur J Surg Oncol 2015;41:1153-61.
66. Guo Z, Zhong Y, Hu B, Jiang JH, Li LQ, Xiang BD. Hepatic resection or transarterial chemoembolization for hepatocellular carcinoma within Milan criteria: a propensity score matching analysis. Medicine 2017;96:e8933.
67. Huang L, Li J, Yan J, Cao J, Liu C, Zhang X, Wu M, Yan Y. Early recurrence after curative resection in oligonodular hepatocellular carcinoma. Hepatogastroenterology 2013;60:28-31.
68. Nojiri K, Tanaka K, Takeda K, Ueda M, Matsuyama R, Taniguchi K, Kumamoto T, Mori R, Endo I. The efficacy of liver resection for multinodular hepatocellular carcinoma. Anticancer Res 2014;34:2421-6.
69. Govindarajan S, Craig JR, Valinluck B. Clonal origin of hepatitis B virus-associated hepatocellular carcinoma. Hum Pathol 1988;19:403-5.
70. Aoki N, Robinson WS. State of hepatitis B viral genomes in cirrhotic and hepatocellular carcinoma nodules. Mol Biol Med 1989;6:395-408.
71. Nomoto S, Kinoshita T, Kato K, Otani S, Kasuya H, Takeda S, Kanazumi N, Sugimoto H, Nakao A. Hypermethylation of multiple genes as clonal markers in multicentric hepatocellular carcinoma. Br J Cancer 2007;97:1260-5.
72. Sirivatanauksorn Y, Sirivatanauksorn V, Bhattacharya S, Davidson BR, Dhillon AP, Kakkar AK, Williamson RC, Lemoine NR. Genomic heterogeneity in synchronous hepatocellular carcinomas. Gut 1999;45:761-5.
73. Nomoto S, Yamashita K, Koshikawa K, Nakao A, Sidransky D. Mitochondrial D-loop mutations as clonal markers in multicentric hepatocellular carcinoma and plasma. Clin Cancer Res 2002;8:481-7.
74. Okuda T, Wakasa K, Kubo S, Hamada T, Fujita M, Enomoto T, Haba T, Hirohashi K, Kinoshita H. Clonal analysis of hepatocellular carcinoma and dysplastic nodule by methylation pattern of X-chromosome-linked human androgen receptor gene. Cancer Letters 2001;164:91-6.
75. Hsu HC, Chiou TJ, Chen JY, Lee CS, Lee PH, Peng SY. Clonality and clonal evolution of hepatocellular carcinoma with multiple nodules. Hepatology 1991;13:923-8.
76. Tsuda H, Oda T, Sakamoto M, Hirohashi S. Different pattern of chromosomal allele loss in multiple hepatocellular carcinomas as evidence of their multifocal origin. Cancer Res 1992;52:1504-9.
77. Hui AM, Kawasaki S, Imamura H, Miyagawa S, Ishii K, Katsuyama T, Makuuchi M. Heterogeneity of DNA content in multiple synchronous hepatocellular carcinomas. Br J Cancer 1997;76:335-9.
78. Furuta M, Ueno M, Fujimoto A, Hayami S, Yasukawa S, Kojima F, Arihiro K, Kawakami Y, Wardell CP, Shiraishi Y, Tanaka H, Nakano K, Maejima K, Sasaki-Oku A, Tokunaga N, Boroevich KA, Abe T, Aikata H, Ohdan H, Gotoh K, Kubo M, Tsunoda T, Miyano S, Chayama K, Yamaue H, Nakagawa H. Whole genome sequencing discriminates hepatocellular carcinoma with intrahepatic metastasis from multi-centric tumors. J Hepatol 2017;66:363-73.
79. Lin DC, Mayakonda A, Dinh HQ, Huang P, Lin L, Liu X, Ding LW, Wang J, Berman BP, Song EW, Yin D, Koeffler HP. Genomic and epigenomic heterogeneity of hepatocellular carcinoma. Cancer Res 2017;77:2255-65.
80. Ng IO, Guan XY, Poon RT, Fan ST, Lee JM. Determination of the molecular relationship between multiple tumour nodules in hepatocellular carcinoma differentiates multicentric origin from intrahepatic metastasis. J Pathol 2003;199:345-53.
81. Hodges KB, Cummings OW, Saxena R, Wang M, Zhang S, Lopez-Beltran A, Montironi R, Nour H, Cheng L. Clonal origin of multifocal hepatocellular carcinoma. Cancer 2010;116:4078-85.
82. Zhao Q, Su CQ, Dong H, Lu XY, Wu MC, Cong WM. Hepatocellular carcinoma and hepatic adenocarcinosarcoma in a patient with hepatitis B virus-related cirrhosis. Semin Liver Dis 2010;30:107-12.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Wang H, Cong WM. New strategy to distinguish clonal origin of RHCC/MHCC between intrahepatic metastasis and multicentric occurrence. Hepatoma Res 2018;4:14. http://dx.doi.org/10.20517/2394-5079.2018.16
AMA Style
Wang H, Cong WM. New strategy to distinguish clonal origin of RHCC/MHCC between intrahepatic metastasis and multicentric occurrence. Hepatoma Research. 2018; 4: 14. http://dx.doi.org/10.20517/2394-5079.2018.16
Chicago/Turabian Style
Wang, Han, Wen-Ming Cong. 2018. "New strategy to distinguish clonal origin of RHCC/MHCC between intrahepatic metastasis and multicentric occurrence" Hepatoma Research. 4: 14. http://dx.doi.org/10.20517/2394-5079.2018.16
ACS Style
Wang, H.; Cong W.M. New strategy to distinguish clonal origin of RHCC/MHCC between intrahepatic metastasis and multicentric occurrence. Hepatoma. Res. 2018, 4, 14. http://dx.doi.org/10.20517/2394-5079.2018.16
About This Article
Special Issue
Copyright
Data & Comments
Data

 Cite This Article 1 clicks
Cite This Article 1 clicks











Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.